Previously we developed the TRUST algorithm 6, 7, 8, utilized to de novo assemble immune receptor repertories directly from tissue or blood RNA-seq data. However, because repertoire sequences from V(D)J recombination and SHM are different from the germline, they are often eliminated in the read-mapping step. Alternatively, RNA-seq data contain expressed TCR and BCR sequences in tissues or peripheral blood mononuclear cells (PBMC). Repertoire sequencing has been increasingly adopted in infectious disease 1, allergy 2, autoimmune 3, tumor immuology 4 and cancer immunotherapy 5 studies, but it is an expensive assay and consumes valuable tissue samples. Following antigen recognition, BCRs also undergo somatic hypermutations (SHMs) to further improve antigen-binding affinity. 10.Both T and B cells can generate diverse receptor (TCR and BCR, respectively) repertoires, through somatic V(D)J recombination, to recognize various external antigens or tumor neoantigens. Practical guidelines for B-cell receptor repertoire sequencing analysis. Deep sequencing and human antibody repertoire analysis. The CAIRR pipeline for submitting standards-compliant B and T cell receptor repertoire sequencing studies to the National Center for Biotechnology Information Repositories. 10.1038/ni.3873īukhari SAC, O'Connor M, Martínez-Romero M, Egyedi A, Debra Willrett D, Graybeal J, et al. Adaptive immune receptor repertoire community recommendations for sharing immune-repertoire sequencing data. Rubelt F, Busse CE, Bukhari SAC, Bürckert J-P, Mariotti-Ferrandiz E, Cowell LG, et al. Reproducibility and reuse of adaptive immune receptor repertoire data. We hope that others will follow suit in the interest of promoting interoperable standards.ĪIRR-seq B cell Rep-Seq T cell antibody immunoglobulin immunology repertoire.īreden F, Luning Prak ET, Peters B, Rubelt F, Schramm CA, Busse CE, et al. Several popular repertoire analysis tools and data repositories already utilize this AIRR-seq data format. It is composed of a tab-delimited format with a specific schema. Our file format emphasizes ease-of-use, accessibility, scalability to large data sets, and a commitment to open and transparent science. In this paper, we describe the work of the AIRR Community's Data Representation Working Group to develop standardized data representations for storing and sharing annotated antibody and T cell receptor data. The AIRR Community has been actively working to standardize protocols, metadata, formats, APIs, and other guidelines to promote open and reproducible studies of the immune repertoire. Increased interest in the immune system's involvement in pathophysiological phenomena coupled with decreased DNA sequencing costs have led to an explosion of antibody and T cell receptor sequencing data collectively termed "adaptive immune receptor repertoire sequencing" (AIRR-seq or Rep-Seq). 13 Department of Genetics and Genomic Sciences and Precision Immunology Institute, Icahn School of Medicine at Mount Sinai, New York, NY, United States.  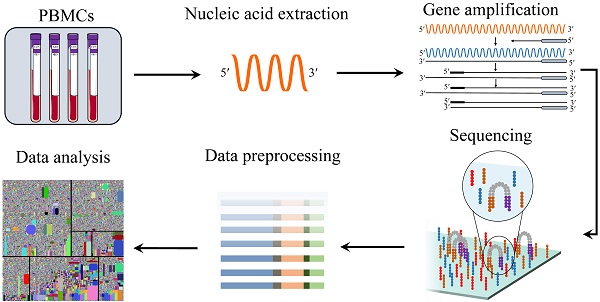
12 Department of Clinical Sciences, UT Southwestern Medical Center, Dallas, TX, United States.11 Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD, United States.10 Fred Hutchinson Cancer Research Center, Seattle, WA, United States.9 Interdepartmental Program in Computational Biology and Bioinformatics, Yale University, New Haven, CT, United States.8 Department of Human Biology, Faculty of Sciences, University of Haifa, Haifa, Israel.7 Department of Microbiology and Immunology, College of Medicine, Drexel University, Philadelphia, PA, United States.6 School of Biomedical Engineering, Science and Health Systems, Drexel University, Philadelphia, PA, United States.5 Department of Biological Sciences, Simon Fraser University, Burnaby, BC, Canada.4 Division of B Cell Immunology, German Cancer Research Center (DKFZ), Heidelberg, Germany.3 Department of Molecular Biology and Biochemistry, Simon Fraser University, Burnaby, BC, Canada.2 Department of Pathology, Yale School of Medicine, New Haven, CT, United States.1 Department of Neurology, Yale School of Medicine, New Haven, CT, United States.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
